Planet recently released support for delivering Planetary Variables to Sentinel Hub. Check out this product announcement and this blog post if you’d like to learn more about what’s been released. To help you explore these new datasets and capabilities, we have also released open datasets for several of our Planetary Variables. We’ve opened these datasets to allow for anybody to explore what this data looks like or to test workflows.
As part of this open data release, data is available for Land Surface Temperature, Crop Biomass, and Soil Water Content. There is 4 years of data from 2019 to 2022 across 4 areas of interest in Uruguay, India, Germany, and the US. You can explore more information about these collections here: https://collections.sentinel-hub.com/tag/planetary-variables/. By clicking through to the specific Planetary Variables, you’ll be taken to their open data pages, such as this Soil Water Content Open Data page.
To make it simple to use these open datasets, they have been hosted in Sentinel Hub. If you don’t already have a Sentinel Hub account, you can sign up here. As part of the 30 day Sentinel Hub trial, you can use the APIs to interact with these open datasets. In addition, if you have a Planet account to order data with, you can also use the 30 day trial to test Sentinel Hub with your own data by using the Sentinel Hub Third Party Data Import API.
If you have any questions about how to use these, feel free to reach out here!
Example Workflow in EO Browser
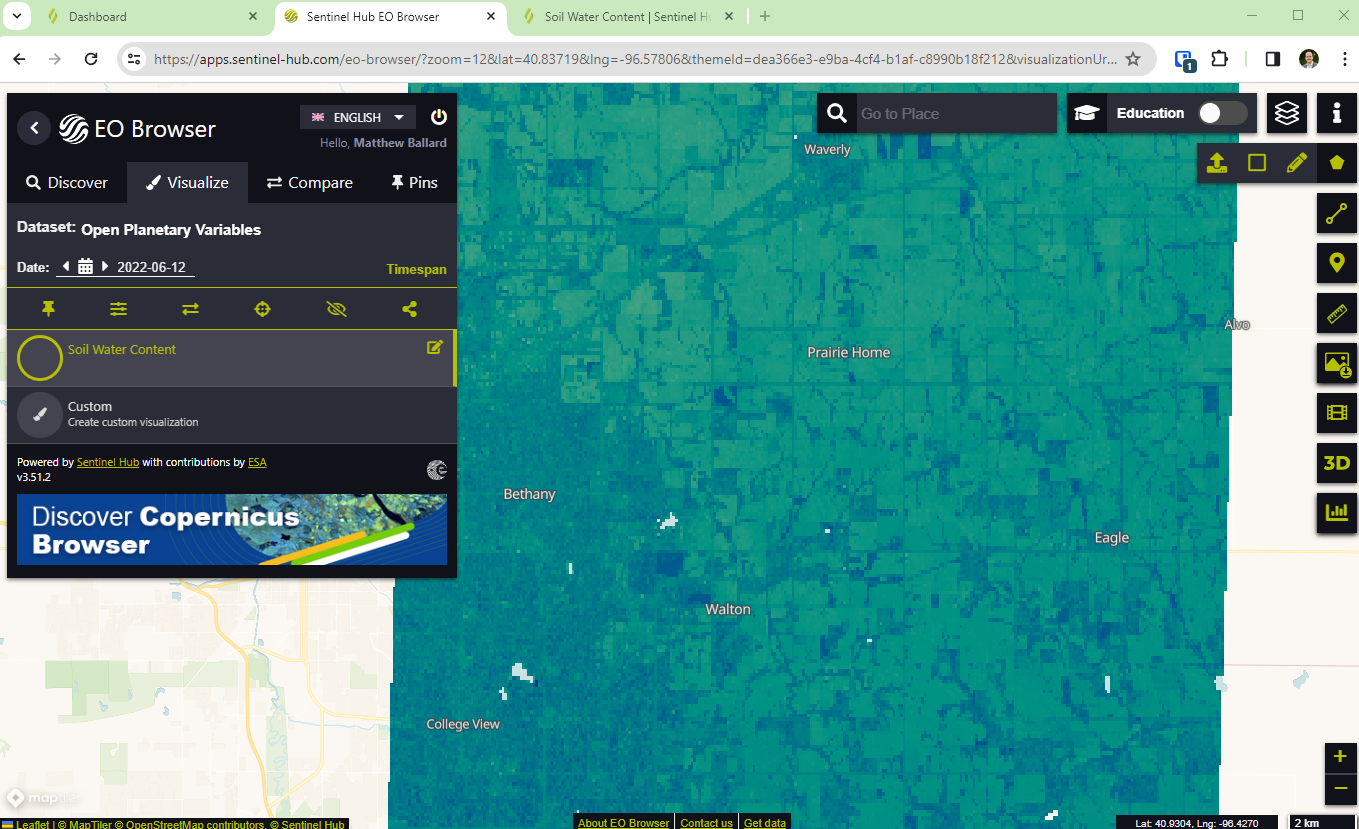
The open datasets can be explored in EO Browser or used with the Sentinel Hub APIs.
To use these datasets with EO Browser, you’ll want to follow these steps:
- Create a new configuration in your Sentinel Hub Dashboard
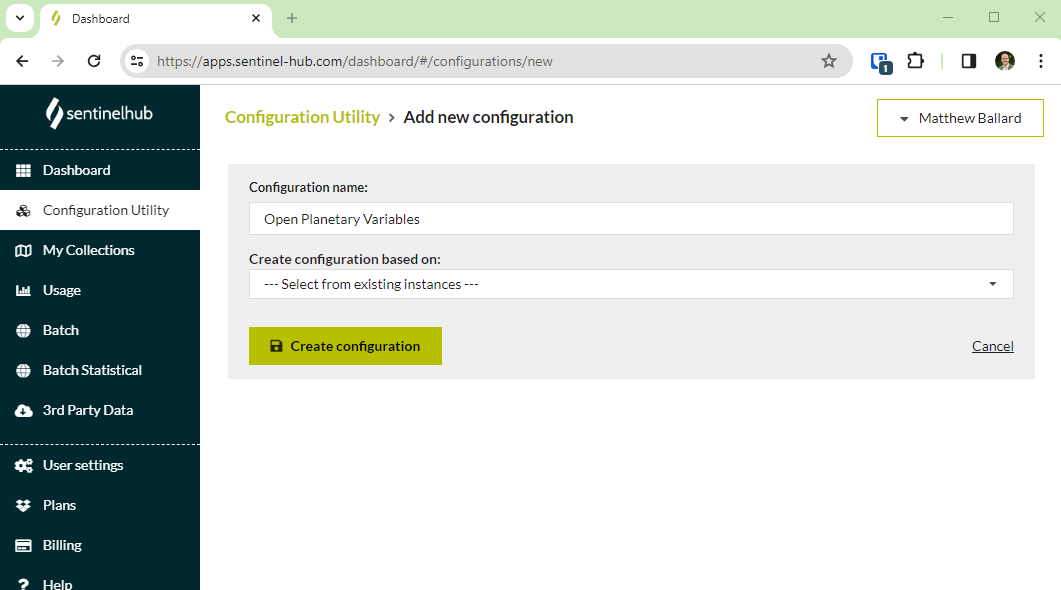
- Add a layer to the configuration and use one of the collection IDs from the open data webpages
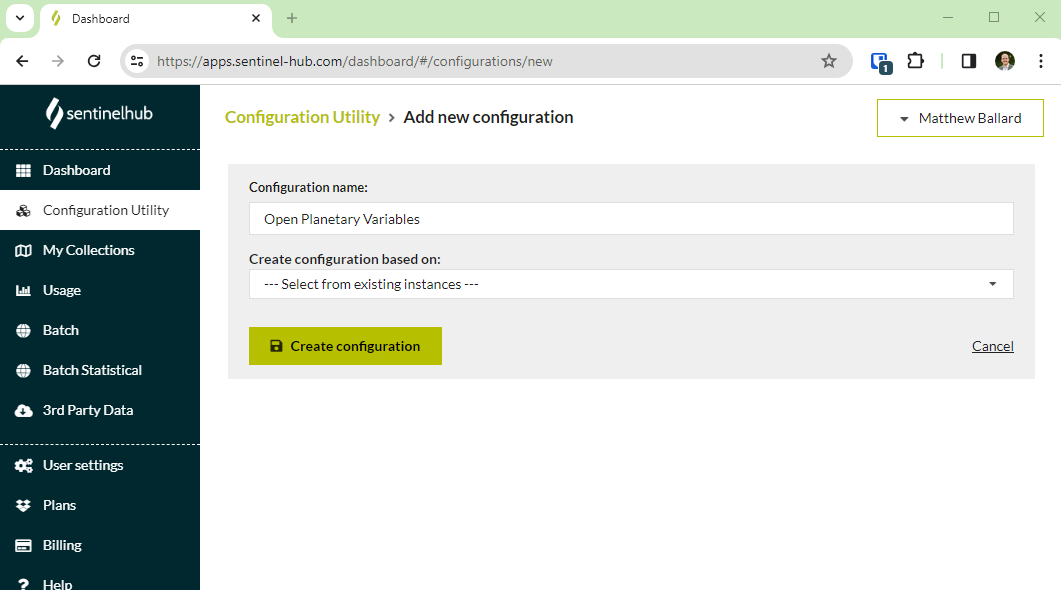
- Add a custom script from our custom scripts repository. This dictates how the Planetary Variables will be visualized. Use one of the scripts that support EO Browser.
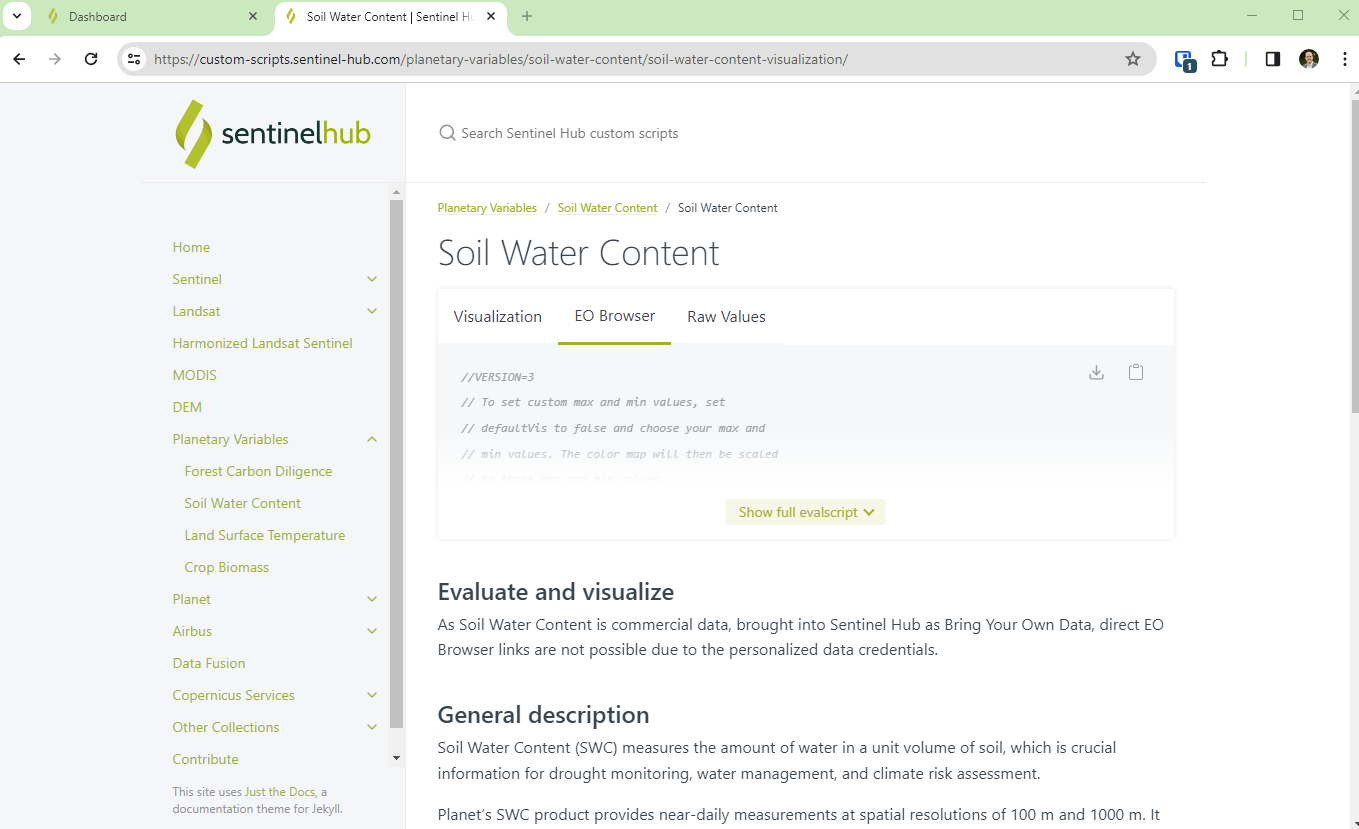
- Save the layer with the collection type set to “BYOC”, the collection ID, and the Data processing set to the custom script.

- From the bottom left, select open in EO Browser. In the top right of EO Browser, add the geojson boundary for one of the areas of interest (found on the open data pages). This will zoom the map to the data extent.
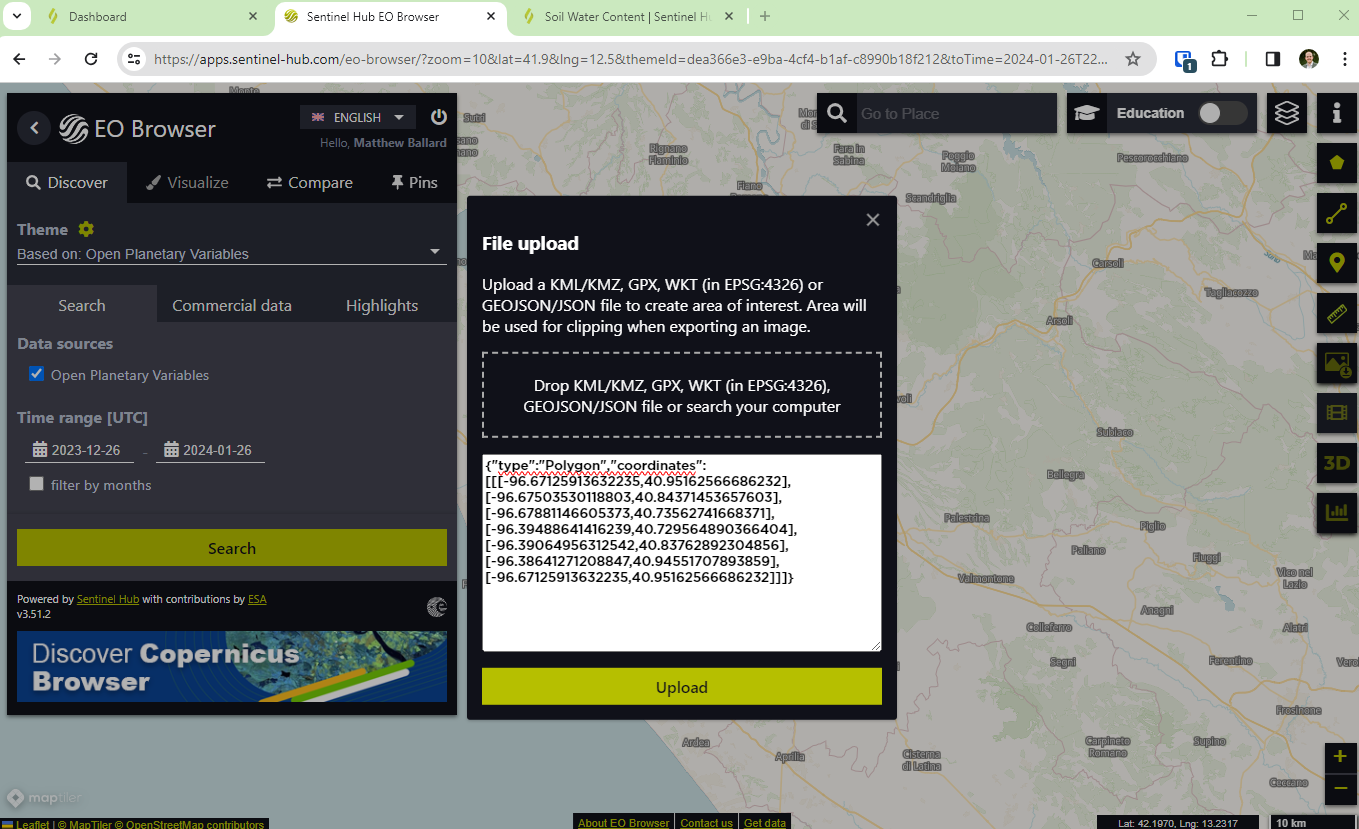
- Change the time range on the left and search for data. Select visualize to view the imagery on the map.